Enhanced CLIP-sequencing (eCLIP)
Transcriptome-wide RNA-binding protein (RBP) mapping and analysis.
Overview
The eCLIP Core provides comprehensive mapping of RNA–protein interactions at transcriptome-wide scale. By utilizing an optimized protocol that decreases requisite amplification by ~1,000-fold, we reduce experimental failure rates and deliver high-complexity libraries for authentic binding site discovery.
Available Services
Maps the precise occupancy of RBPs on RNA to deduce biological function and target transcript regulation.
Measures ribosome occupancy to distinguish mRNA targets of ribosomes with varying protein compositions, offering mechanistic insight into translation efficiency.
Simultaneously captures up to 10 RBPs from a single sample. Ideal for establishing rapid, highly parallelized RBP–RNA networks from limited clinical inputs.
Bioinformatics Pipeline & Data Delivery
All eCLIP sequencing data is rigorously processed to identify high-confidence, reproducible RNA-binding windows using our standardized SKIPPER pipeline. Results are securely delivered via Globus, providing researchers with reliable, high-speed data transfer directly to their institutional or cloud endpoints.
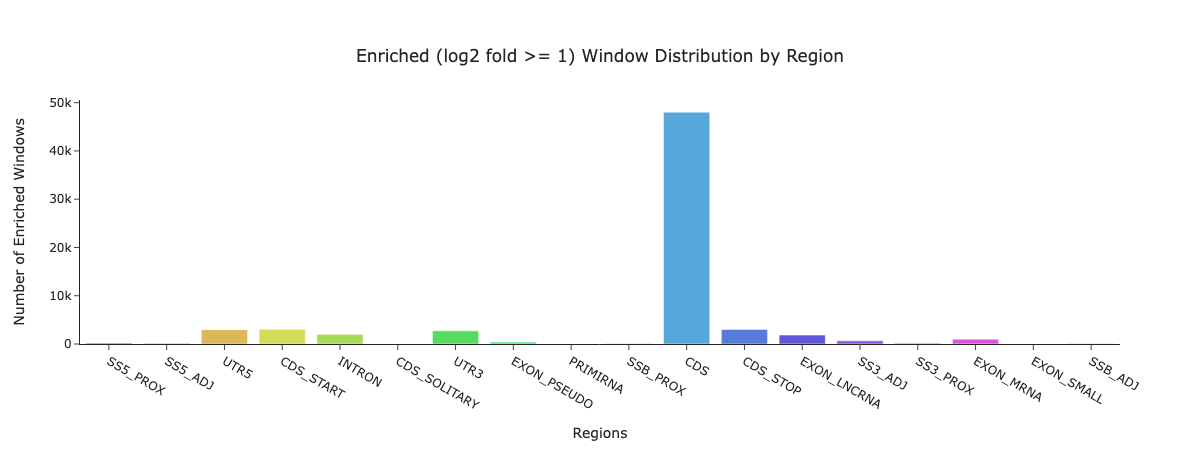
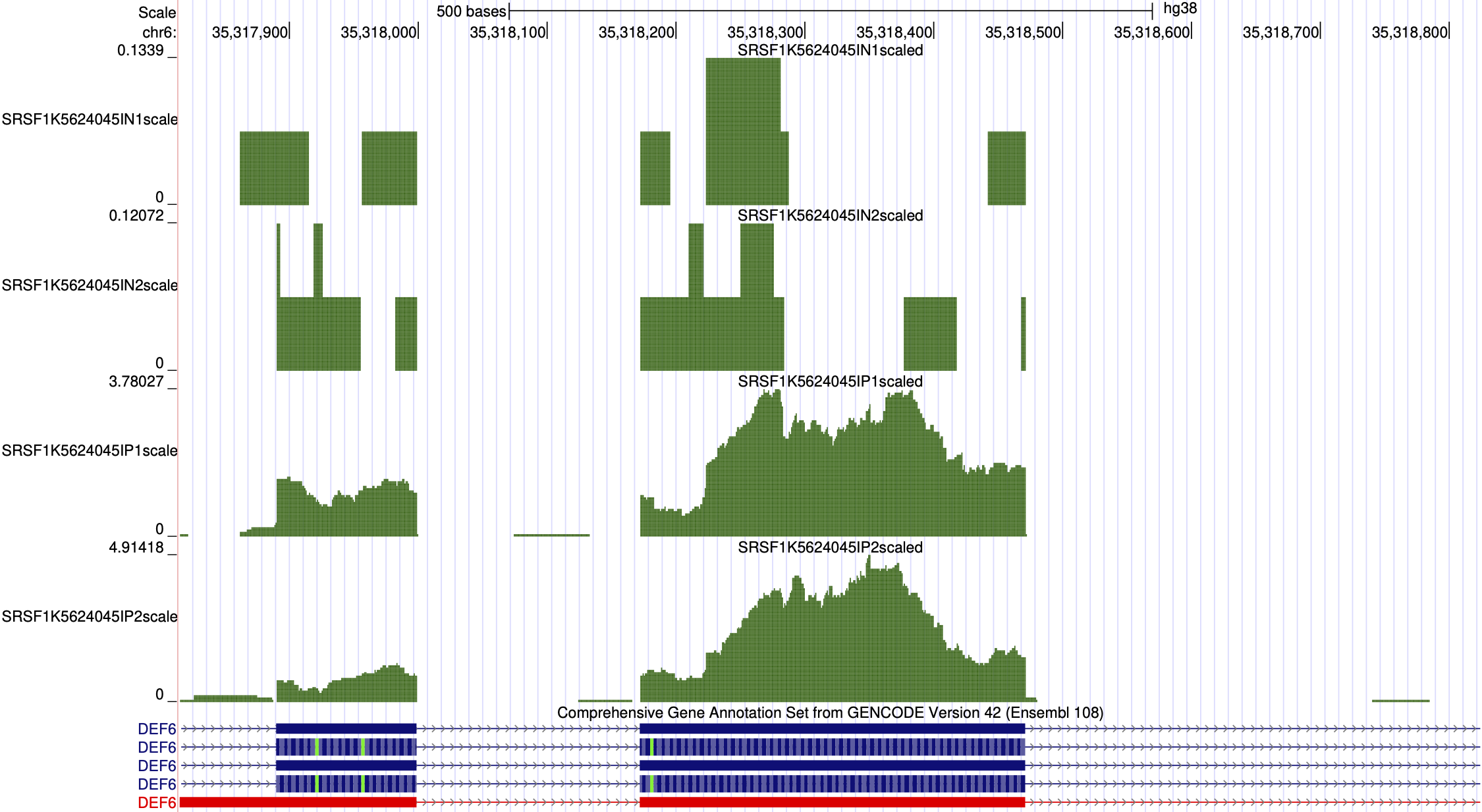
Upon project completion, your data is loaded into our interactive hub. Researchers receive comprehensive quality control metrics and analysis figures, including region distributions, basic read statistics, and reproducible enriched windows autolinked to UCSC's genome browser.
| Sample | Initial Reads | Trimmed Reads | Deduped Reads | Usable % |
|---|---|---|---|---|
| SRSF1_K562_4045_IP_1 | 25,141,300 | 24,927,766 | 17,194,358 | 0.68 |
| SRSF1_K562_4045_IN_1 | 20,572,437 | 20,541,064 | 14,936,346 | 0.73 |
https://g-ba9d59.0ed28.75bc.data.globus.org/analysis/public/eclip_example/
# UCSC Genome Browser Trackhub — paste into My Data › Trackhubs › Connected Hubs:
http://s3-us-west-2.amazonaws.com/yeolab-trackhubs/20240722-ExampleeCLIP-SKIPPER/20240722-ExampleeCLIP-SKIPPER.hub.txt
How It Works
eCLIP captures direct, in vivo RNA–protein interactions through UV crosslinking followed by immunoprecipitation and strand-specific library preparation. UV irradiation induces covalent bonds between RBPs and their directly contacted RNA targets, after which stringent purification and a size-matched input (SMInput) control enable precise, quantitative identification of reproducible binding windows across the transcriptome.

- 1. Van Nostrand EL, et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat Methods. 2016;13(6):508–514. PMID: 27018577
- 2. Van Nostrand EL, et al. Transcriptome-wide identification of RNA-binding protein binding sites using seCLIP-seq. Nat Protoc. 2022;17(5):1223–1265. PMID: 35322209