Circular RNA sequencing (circRNA-seq) Core
High-sensitivity biomarker discovery and complete topological sequencing.
Overview
The circRNA-seq Core specializes in the extraction, enrichment, and sequencing of covalently closed RNA molecules. Our proprietary library preparation methods allow for robust profiling of clinical specimens and FACS-isolated cells, yielding highly sensitive data for the study of microRNA sponges, RBP decoys, and novel therapeutic vectors.
Available Services
A highly sensitive assay requiring orders of magnitude less input than published standard methods. Designed specifically for profiling rare cells and precious clinical samples.
Input Requirement: As low as 50 ng total RNAUtilizes Nanopore instrumentation to sequence the entire RNA circle. By bypassing the bioinformatic limitations of short-read backsplicing detection, this assay perfectly preserves alternative splicing information and regulatory mechanisms.
Customized pipelines to study trans-factor interactions, specifically mapping RBP-circRNA binding kinetics and evaluating circRNA translation efficiency.
Bioinformatics Pipeline & Data Delivery
All circRNA sequencing data is rigorously processed to identify high-confidence backsplicing junctions and quantify circular RNA expression using a standard, up-to-date circRNA detection pipeline. Results are securely delivered via Globus, providing researchers with reliable, high-speed data transfer directly to their institutional or cloud endpoints.
Upon project completion, your data is loaded into our interactive hub. Researchers receive comprehensive quality control metrics, backsplice junction counts, and navigable data tables:
IExample output: identified circRNA Positions with confidence score.
| seqname | start | end | gene_name | gene_type | circ_type | score | strand |
|---|---|---|---|---|---|---|---|
| chr1 | 805799 | 810170 | AL669831.3 | transcribed_processed_pseudo... | exon | 0.1758 | - |
| chr1 | 1217045 | 1228641 | SDF4 | protein_coding | exon | 0.1172 | - |
| chr1 | 1223244 | 1223968 | SDF4 | protein_coding | exon | 1.758 | - |
| chr1 | 1256045 | 1267992 | UBE2J2 | protein_coding | exon | 0.0586 | - |
| chr1 | 1477849 | 1478627 | ATAD3B | protein_coding | intron | 0.5274 | + |
Example output: Identifying and comparing distribution of circRNA among six submitted samples.
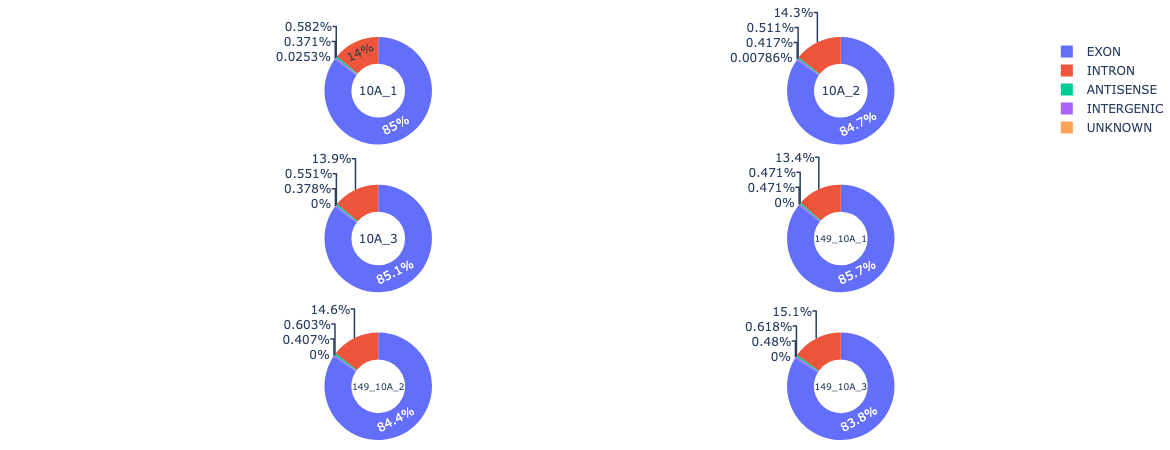
Acknowledgment Policy
Standard Language: "We thank the UC San Diego Center for RNA Technologies and Therapeutics for circRNA library preparation and sequencing."