The RNA Interactome Toolkit
From highly multiplexed binding profiles to functional RNA drug screens, we provide the cutting-edge wet-lab assays and robust data structures needed for the next frontier of RNA biology.
Enhanced CLIP-seq (eCLIP)
Map RNA-protein interactions transcriptome-wide at single-nucleotide resolution. Includes standard eCLIP, Ribo-eCLIP, and highly multiplexed OligoCLIP for limiting samples.
Explore eCLIP CoreCircular RNA (circRNA-seq)
Empower biomarker discovery and functional biology research with highly sensitive detection (50 ng input) and full-circle Nanopore long-read resolution.
Explore circRNA CoreCustom RNA Synthesis
Precision in vitro transcription (IVT) featuring custom template design, in vivo modifications, and high-throughput automated synthesis via the Telesis BioxP.
Explore Synthesis CoreSeamless Computational Integration
Our sequencing cores don't just hand over raw FASTQ files; we provide structured, high-fidelity matrices engineered for modern computational labs. All data is transferred securely via Globus.
- Immediately interpretable results: Processed datasets are automatically integrated into our data portal, allowing users to explore, visualize, and interpret primary results without additional setup.
- Human and machine readable outputs: Text/TSV-formatted outputs designed for immediate ingestion into standard computational workflows and office environments.
- State-of-the-art analysis: Data is processed using scalable, reproducible workflows maintained in-house. Our pipelines are built on open-source tools, transparently versioned, and continuously updated to incorporate the latest methodological advances in computational genomics.
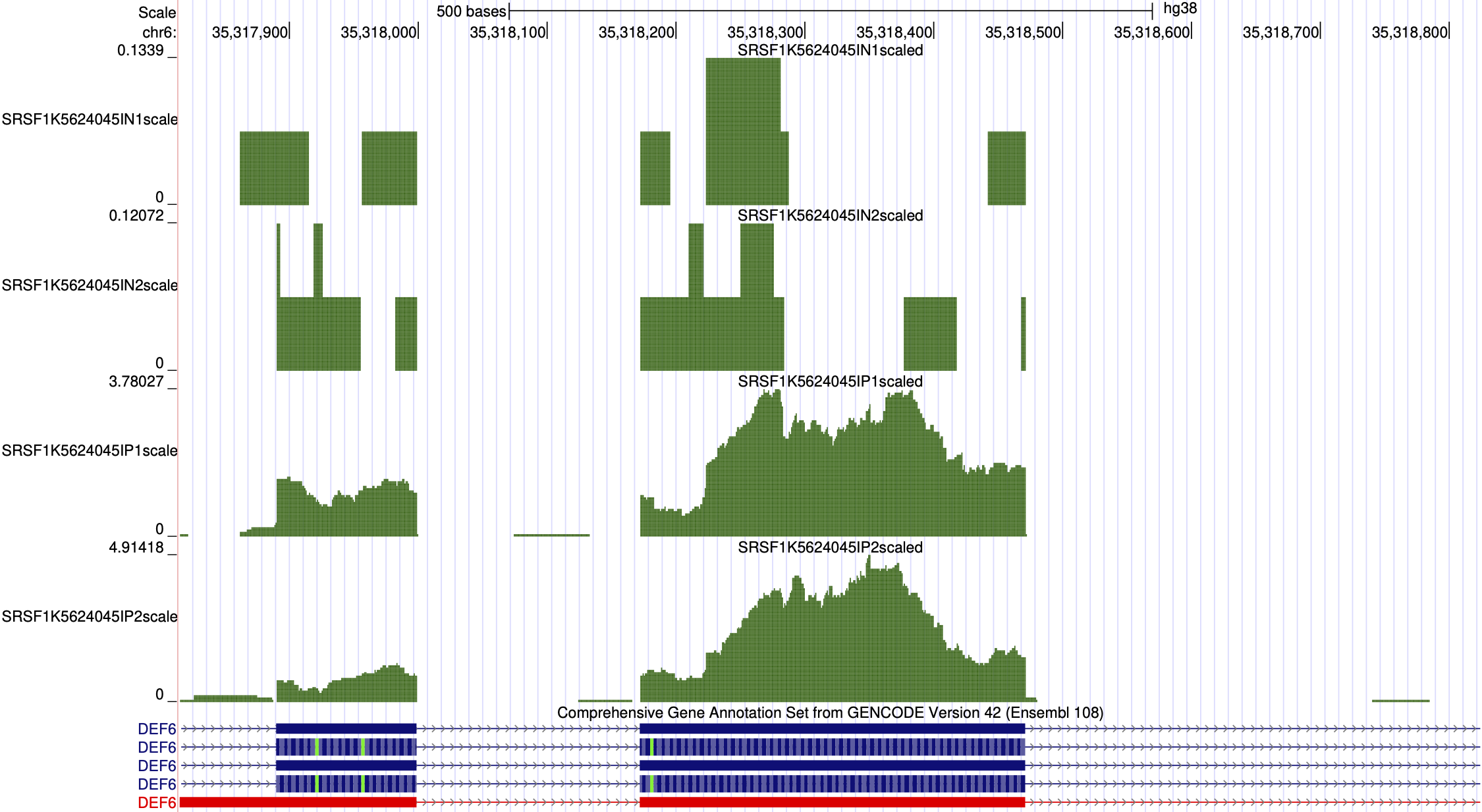
Project Workflow
From experimental design to data handoff, our streamlined process ensures rigorous quality control.
Initiation
Discuss experimental design, custom goals, and cost estimates.
Preparation
Prepare cell lines/tissue. Conduct experimental validation.
Submission
Submit required manifest and ship/drop-off physical samples.
Processing
Protocol execution with primary data analysis and QC.
Data Handoff
Secure transfer of raw sequencing files and optional bioinformatic analysis.
- Initiation: Discuss design and goals.
- Preparation: Prepare samples & experimental validation.
- Submission: Submit manifest and samples.
- Processing: Sequencing, QC, and primary analysis.
- Data Handoff: Transfer of data and analysis.